Gene Information
This web page was produced as an assignment for Gen677 at UW-Madison Spring 2009
Phylogenetic tree
This is an analysis using the DNA sequence to look at the relatedness of the SIRT6 gene in model organisms
DNA motif
This is an analysis of the SIRT6 human DNA sequence to look for possible binding site for transcription factors
DNA Alignment
This is an analysis looking at the simularity base pair simularity of the SIRT6 gene in different model organisms
Microarray
This is an analysis that looks at different expression charatistics of SIRT6 under different conditions
Gene Ontology
This is an analysis for specific terms that relate to SIRT6 function, location and cellular component
SIRT6 gene
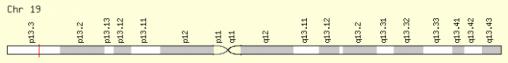
The SIRT6 gene in human is located on chrosome 19, the gene is a 1638 base pair gene. Also in humans there are four isoforms according to uniport, but little is known about these isoforms and ther functions. I will mainly focus on the human form of SIRT6 because I would like to look at how this gene compares to other organism and see if model organism can closly simulate what this gene might do in humans.
Homologs of SIRT6 gene
SIRT6 Mus musculus DNA sequence
NM:181586.2
Protein E value: 3e-174
DNA E value: 0
SIRT6 Danio rerio DNA sequence
NM: 001002071.1
Protein E value: 3e-128
DNA E value: 3e-102
SIRT6 Gallus gallus DNA sequence
NM: 001039320.1
Protein E value: 3e-148
DNA E value: 0
SIRT6 Rattus norvegicus DNA sequence
NM: 001031649.1
Protein E value: 7e-173
DNA E value: 0
SIRT6 Pan troglodytes DNA sequence
XM: 001138012.1, XM: 001137928.1, XM: 001137581.1
Protein E value: 0
DNA E value: 0
SIRT6 Drosophila melanogaster DNA sequence
NM: 141733.1
Proein E value: 7e-88
DNA E value: 4e-4
SIRT6 Homo sapiens DNA sequence
NM: 016539.1
All the E values where obtained by comparing the different DNA or protein sequences against the human sequence of SIRT6. As you can see the Gallus gallus and Pan troglodytes protein sequence is very similar to the human form. One caveat of the blast analysis comparing DNA sequence was that I could only do some what similar search and some of the sequences where only covered upto 61% of the sequence to the human DNA sequence. So this means that most of the sequence wasnt similar to the human and only the parts that were somewhat similar were compared so this may have made for the higher E values.
Mark Devries
Email [email protected]
last updated 1/22/09
http://www.gen677.weebly.com
